Translate your Bio-attribution Research into National Security Impact
Register for Virtual Competition to Win a Share of $180,000 in Prizes
In an era of unprecedented biological data generation, the ability to rapidly determine the origin of a biological event — whether natural, accidental, or intentional — is a critical component of national security and public health. To meet the challenge of finding the "needle in a haystack" within this data deluge, the Defense Advanced Research Projects Agency (DARPA) has launched the Bio-Attribution Challenge.
This virtual competition calls on innovators to develop a new generation of tools capable of analyzing petabyte-scale datasets in near real-time, far exceeding the capacity of current systems. The goal is to revolutionize how we identify and trace the source of biological sequences, ensuring a faster, more effective response to potential threats.
"The ability to rapidly and accurately identify the source of a biological sequence, whether natural or engineered, is a critical national security capability," said Abhishek Singharoy, Ph.D., program manager for the Bio Attribution Challenge. “We’re calling on creative researchers to help us catalyze a new generation of tools that can find the proverbial 'needle in a haystack' within data environments of unprecedented scale and complexity."
The competition is structured in two rounds:
Round 1: Detection | 2 months – Focuses on the accurate identification and characterization of pathogens within complex environmental samples.
Round 2: Attribution | 1.5 months – Challenges participants to determine the origin of engineered pathogens by identifying unique physical, chemical, or design signatures.
A total of $180,000 in monetary prizes will be awarded to the top three performers in each round:
| Round 1 | |
| First place | $50,000 |
| Second place | $30,000 |
| Third place | $10,000 |
| Round 2 | |
| First place | $50,000 |
| Second place | $30,000 |
| Third place | $10,000 |
In addition to monetary prizes, the Bio Attribution Challenge offers significant opportunities for participants, including:
- The chance to present their work to leaders from various branches of the U.S. Department of War
- A platform to learn from and collaborate with other leaders and partners from industry, government, and academia
- Consideration for follow-on opportunities such as Other Transaction Agreements (OTAs) and Cooperative Research and Development Agreements (CRADAs)
- Recognition for outstanding innovation, data efficiency, and other significant achievements, including an overall "Best in Show" award
The competition will culminate in an awards ceremony on June 30, 2026.

Round 1 Winners
The first round of the DARPA Bio-Attribution Challenge, a rigorous two-month competition focused on the detection and characterization of complex biological sequences, has concluded.
The challenge now progresses to its next phase.
Invitations have been sent to qualifying participants for Round 2, which will focus on determining the attribution and origins of engineered biological events.

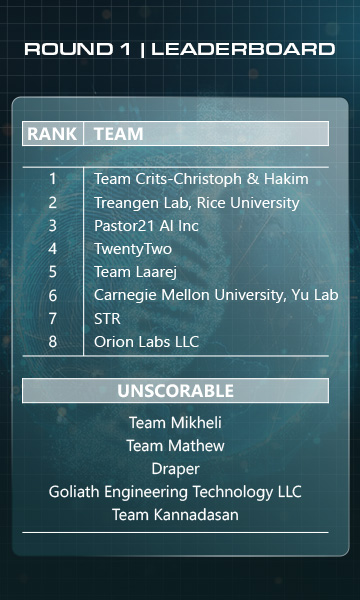
These are the top-performing teams in the "Detection" phase of biological sequence analysis.
A Purely Computational Challenge
This is a purely computational challenge; no actual pathogens or biological materials will be used. All data is deliberately curated and developed by Lawrence Livermore National Laboratory (LLNL) to mimic realistic and complex scenarios without disclosing sensitive information. Participant software will be run in a secure, government-controlled environment.
DARPA encourages participation from varied teams and individuals with expertise in bioinformatics, data science, high-performance computing, information theory, and machine learning.

