Successes in first two years of Safe Genes program establish technological foundations and ground truth in support of DARPA’s emerging, adaptable resources for secure genome editing research
Oct. 15, 2019
DARPA launched the Safe Genes program in 2017 to establish a “safety by design” strategy for guiding the development of an array of powerful, emergent genome editing technologies. Consistent with the National Biodefense Strategy published last year, DARPA’s goals for Safe Genes are to mitigate the risks and security concerns related to the accidental or intentional misuse of such technologies and, at the same time, enable the pursuit of novel genetic solutions that support public health and military force protection and readiness.
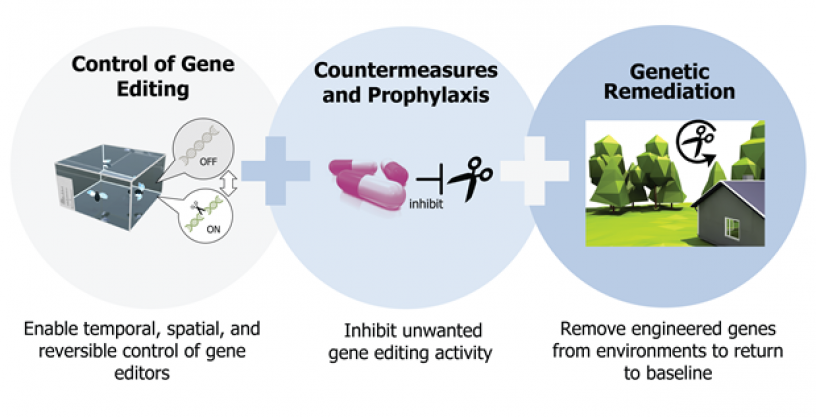
In the first two years of the program, Safe Genes performers worked to gain a fundamental understanding of how genome editing technologies function and how they might be controlled or countered. To do so, the researchers developed new molecular tools and measurements and led discovery through laboratory experiments and computer simulations. Encouraging results from this first phase of the program support the ongoing development of safety mechanisms and countermeasures to control, block, and reverse the effects of genome editors, as well as the creation of new technologies to safely, responsibly, and predictably apply them when warranted.
“During the first phase of Safe Genes, we focused on ground truth, technological foundations, and early proofs of concept to determine which research pathways show the most promise,” said Renee Wegrzyn, the Safe Genes program manager. “The remaining Safe Genes teams are spearheading unique approaches that support development of a layered, modular, adaptable solution set.”
For example, the research team led by Massachusetts General Hospital (MGH) is pursuing a multifaceted approach to improving gene editing capabilities through the introduction of greater control over when and where in the genome that editors function — necessary advances for the use of editors for therapeutic purposes. Already they have developed tools for accurately measuring unintended edits known as off-target effects, along with open-source software to rapidly analyze sequencing data, and they have applied these technologies to the informed design of novel guides and editors that improve overall performance while reducing off-target effects.
The MGH team first demonstrated its design technology by measuring and validating the precision of the new editors during laboratory experiments in mouse models.i With the technology, known as VIVO for “Verification of In Vivo Off-targets,” the team was able to identify with great sensitivity any off-target mutations following introduction of a CRISPR-Cas genome editor.
The MGH team subsequently contributed to the research community an open-source software resource called “CRISPResso2” for rapid analysis of genome editing sequences.ii Prior to CRISPResso2, the long processing times and inaccuracies associated with earlier analysis tools made the study of presumed off-target effects largely infeasible. With CRISPResso2, the results of an experiment using modern high-throughput sequencing technologies can now be processed and analyzed in less than one minute with a negligible false-positive rate (<0.01).
MGH also delivered new findings and tools to improve the performance of base editors, a class of gene editing proteins that scientists initially thought might avoid some off-target effects because they function without the need to make a full break in DNA structure. However, the MGH team reported that even base editors can contribute to off-target effects by unexpectedly editing RNA transcripts.iii,iv The team discovered that base editors are capable of both editing mRNAs across the entire transcriptome and self-editing their own transcripts, potentially leading to the expression of alternative proteins. MGH addressed this problem by using structure-guided engineering to create base editors that induce substantially fewer RNA off-target alterations in human cells and lower self-editing to undetectable levels while still preserving, and even improving, desired DNA-editing activity. The research is an important first step toward translating CRISPR-Cas genome editing tools from laboratory use into clinical use.
Other Safe Genes researchers are developing tools to counter genome editing activity and protect the genome integrity of organisms against unwanted changes. Teams led by the University of California, Berkeley (UCB), and MGH have characterized the ability of natural, anti-CRISPR proteins to block changes that would otherwise be induced by genome editors. Their work encompasses not only proteins effective against Cas9, but also, for the first time, anti-CRISPR-Cas12a proteins demonstrated to block or diminish edits in human cell cultures. In a series of four analyses, the researchers set the groundwork for understanding how to protect against the diversity of current and future CRISPR genome editors.v,vi,vii,viii
Complementing this anti-CRISPR protein work, the Safe Genes team led by the Broad Institute/Brigham and Women’s Hospital/Harvard Medical School (Broad) also developed high-throughput methods for detection of Cas proteinsix and inhibitors.x Using these methods, the researchers discovered new small-molecule inhibitors and activators, and applied them to achieve spatiotemporal, light-activated control of Cas9.xi Going forward, these technologies could be applied, for example, to increase the precision of genome editing activity in the body during therapeutic applications of editors or to deny the effects of unsanctioned use of editors against an individual or in a given environmental setting.
At the macro level, Safe Genes teams are studying control mechanisms for and countermeasures against gene drives — genome editing tools designed to quickly push genetic modifications into large numbers of a particular species within a given area. Gene drives have been discussed in public health circles as a potential solution for containing the spread of infectious diseases such as malaria and Zika by controlling the ability of insect vectors to transmit pathogens. However, at the time DARPA launched Safe Genes, existing research had not adequately developed strategies to limit the spread of gene drives in a predictable manner, and had not accounted for the numerous environmental and genetic factors that would introduce complexity were a gene drive released into the real world. That gap in knowledge not only impeded forward progress in developing disease control strategies, but also served as a blind spot in national security preparedness.
Safe Genes set out to establish the ground truth of gene drive performance. Program performers are blending mathematical modeling, cage trials, and advanced experiments conducted in biosecure, simulated natural environments to add layers of complexity, gain an understanding of how gene drive technologies might actually function, and inform development of tools that can control, counter, or reverse their effects. Safe Genes teams are also defining how various failure modes, including functional genetic resistance,xii may prevent a drive from taking hold.
Safe Genes researchers speak publicly about their ongoing efforts while they await publication of initial results. Dr. Omar Akbari, principal investigator for the team led by University of California, Riverside (UCR), and University of California, San Diego, focuses on engineering self-limiting gene drives that must meet certain threshold requirements before they can propagate through a population.xiii Dr. Amit Choudhary, principal investigator for the Broad team, is developing means of engineering chemical dependencies into Cas9 to tune the activity of gene drives.xiv
While research is ongoing to understand gene drive performance and improve controls, the Safe Genes program has also produced a next-generation, CRISPR-based vector-control technology that is scalable, self-limiting, highly effective, and safe. The UCR-led team published details of a novel sterile-insect technique that has the potential to prevent the spread of diseases such as dengue and Zika by locally reducing vector numbers in a predictable and controllable manner.xv Early and positive results like these from the program are creating space for new investments by other government agencies, non-governmental organizations, and universities.
“What I find most inspiring about the Safe Genes results to date is that our work flips the traditional understanding of dual-use research on its head,” Wegrzyn said. “We set out to protect against misuse of genome editors, and by virtue of making progress in that mission, we’re also laying the groundwork for safe, predictable, and potentially transformative applications of the technology to preserve the health of service members and support public health more broadly. It’s a good example of how DARPA can lead the way in an emerging field to facilitate other communities and organizations digging deeper.”
The Safe Genes teams are led by universities and university-affiliated medical research centers conducting contracted fundamental research. Recently, the Safe Genes program transitioned into its second phase with four teams continuing: Broad, MGH, UCB, and UCR. These teams will continue to publish results as they become available. Additionally, DARPA encourages the teams to adjust their projects as necessary to incorporate new findings and technologies from elsewhere in the rapidly evolving field of genome editing.
i https://www.nature.com/articles/s41586-018-0500-9
ii https://www.nature.com/articles/s41587-019-0032-3
iii https://www.nature.com/articles/s41586-019-1161-z
iv https://www.nature.com/articles/s41587-019-0236-6
v http://science.sciencemag.org/content/early/2018/09/05/science.aau5138
vi http://science.sciencemag.org/content/early/2018/09/05/science.aau5174
vii https://www.nature.com/articles/s41594-019-0208-z
viii https://www.cell.com/molecular-cell/abstract/S1097-2765(17)30935-8
ix https://pubs.rsc.org/en/content/articlehtml/2019/sc/c8sc03426e
x https://www.cell.com/cell/fulltext/S0092-8674(19)30395-2
xi https://onlinelibrary.wiley.com/doi/abs/10.1002/anie.201900788
xii https://www.nature.com/articles/nbt.4245
xiii https://www.genengnews.com/insights/crispr-accelerated-gene-drives-pump-the-brakes/
xiv https://www.genengnews.com/insights/a-toolbox-for-keeping-crispr-in-check/
xv https://www.nature.com/articles/s41467-018-07964-7
# # #
Media with inquiries should contact DARPA Public Affairs at outreach@darpa.mil
Associated images posted on www.darpa.mil and video posted at www.youtube.com/darpatv may be reused according to the terms of the DARPA User Agreement, available here: http://go.usa.gov/cuTXR.
Tweet @darpa
